Transverse program
The objective of this program is to encourage the association of several teams to answer a common scientific question, which could not be addressed individually by the teams, and would therefore be approached in an interdisciplinary and collaborative way. A large proportion of the Labex renewal budget, € 3 million, is dedicated to this program.
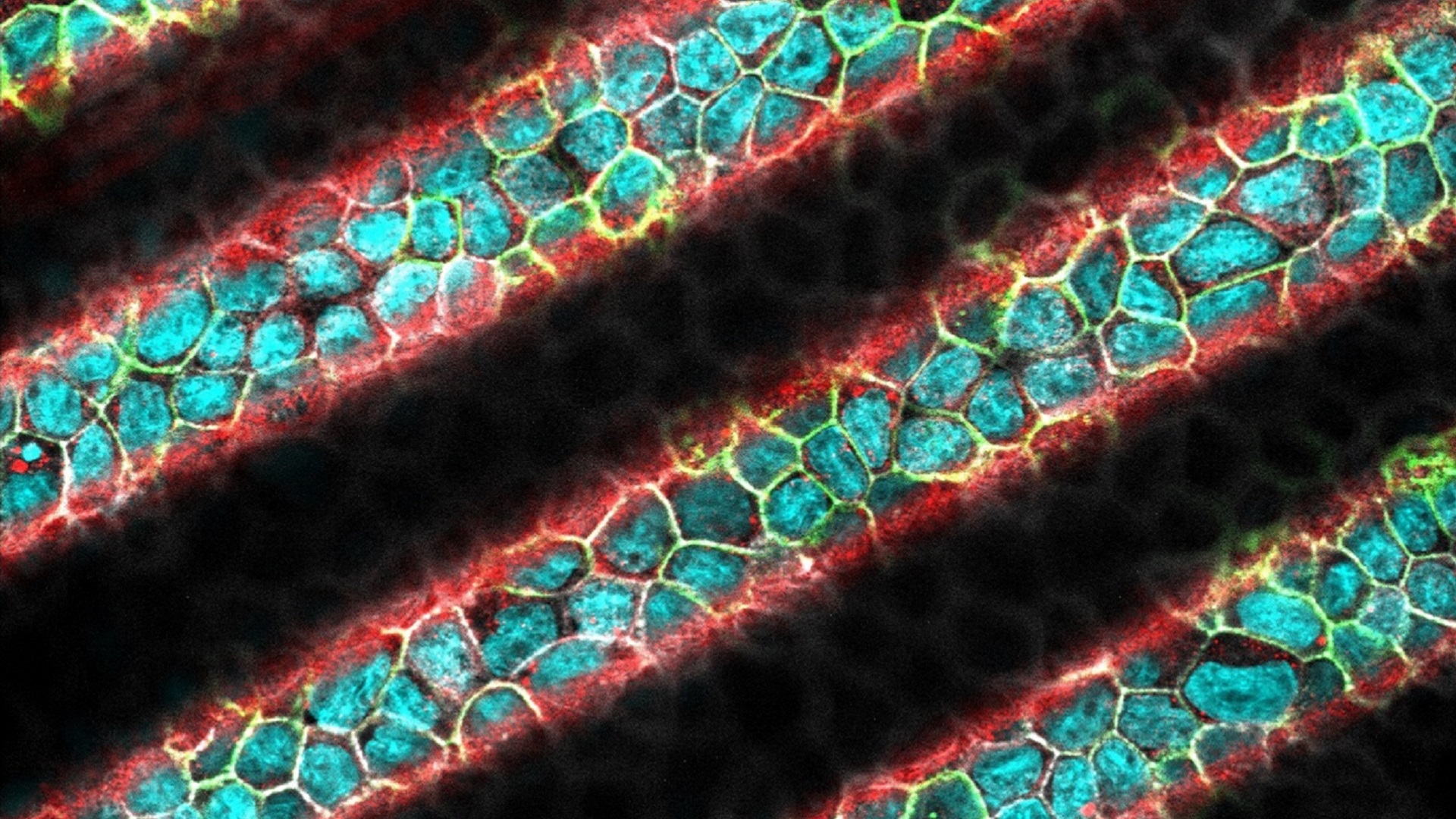
Sinusoidal epithelium
MDCK epithelium on a substrate with a sinusoidal profile; nuclei appear in cyan, Ecadherins in green, apical membrane in red and F-actin in white.
© Sylvie HENON, Matter and Complex Systems laboratory
The Transverse program is divided into three phases.
First phase
Two projects were selected and launched in September 2020:
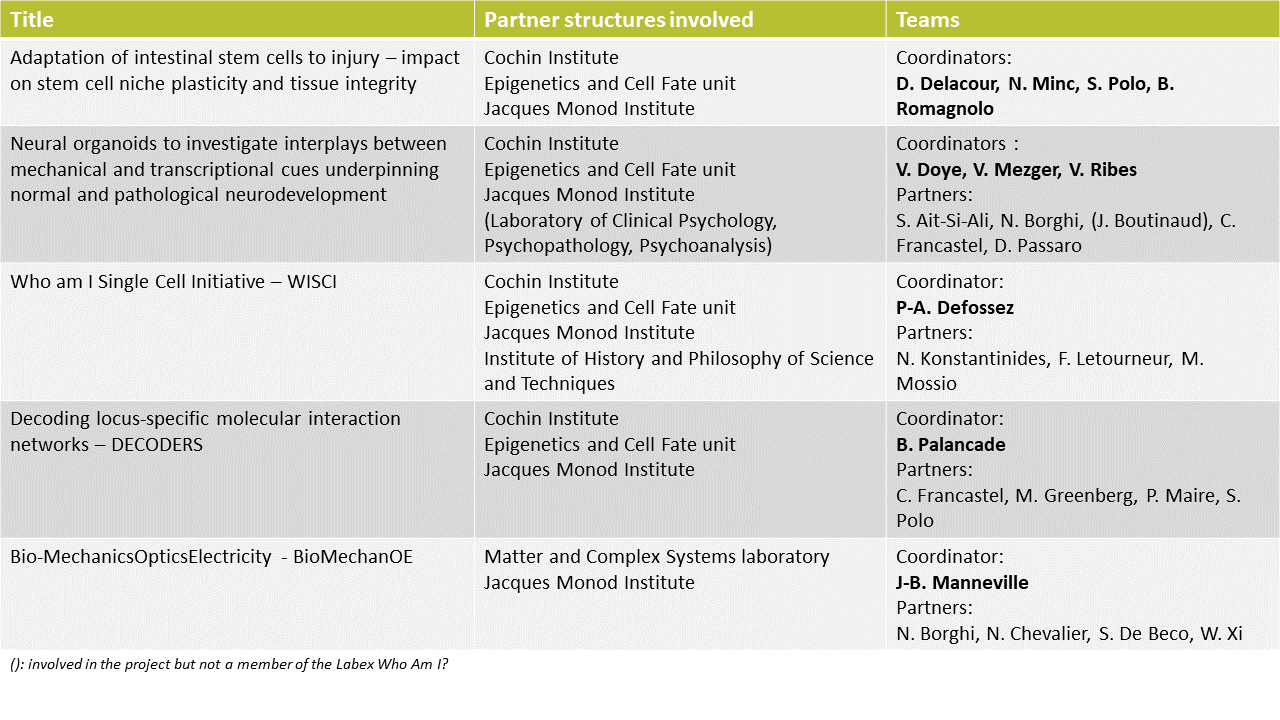
1) Adaptation of intestinal stem cells to injury – impact on stem cell niche plasticity and tissue integrity
In this project, the question of the regulation of the fate of intestinal stem cells is addressed using intestinal organoids as a model system. The objectives of this work, bringing together four teams, are to define how cell division contributes to determining the fate of intestinal stem cells and how their identity is maintained.
2) Neural organoids to investigate interplays between mechanical and transcriptional cues underpinning normal and pathological neurodevelopment
In this project, eight teams with complementary expertise aim to study the interactions between mechanical and transcriptional signals that dictate the spatio-temporal dynamics of fate and cellular form, underlying the normal or pathological development of the human central nervous system.
Second phase
Two projects were selected and launched in May 2021:
1) Who am I Single Cell Initiative – WISCI
This project brings together four teams whose objective is to remove the limitations to the performance of Single-Cell genomics experiments at Université Paris Cité. To do this, this project builds on and strengthens existing platforms. In addition, interactions between biologists, philosophers and historians of science are promoted around the study of the concept of the cell.
2) Decoding locus-specific molecular interaction networks – DECODERS
Bringing together teams from three partner structures, this consortium aims to decode locus-specific molecular interaction networks. The development of spatially resolved proteomics and transcriptomics approaches allows this study at the level of individual genomic sequences.
Third phase
One project was selected and launched in December 2021:
Bio-MechanicsOpticsElectricity – BioMechanOE
This last selected project has two main objectives: combine several mechanobiological and optical techniques on the same set-ups to make simultaneous measurements, and use these technologies to explore the links between tumor development and metabolism. Led by two main Labex’s partners, it combines the expertise from four structures and includes a science outreach part.
À lire aussi

Closing Conference 2024 – Interdisciplinary perspectives on identity
The Labex Who Am I? invites you to its closing conference, “Interdisciplinary Perspectives on Identity: from Single Cells to Organism and Beyond”, which will take place on November 27-29 in Paris. The Labex Who Am I? final conference will take place on November 27, 28...

Pint of Science festival 2024
Once again this year, the Labex Who Am I? is partnering with the Faculty of Sciences of the Université Paris Cité, the Genetics and Epigenetics New Education (G.E.N.E.) Graduate School, and the Major Research and Innovation Domain BioConvergence for Health...

Visiting Professors programme: Matthias Peter
As part of its Visiting Professors programme, the Labex Who Am I? is delighted to welcome professor Matthias Peter, professor of Biochemistry and group leader at ETH Zurich, Switzerland. © Peter group, ETH Zurich Matthias Peter...

Rhythms and resonances: coming into contact with otherness
The Labex Who Am I? co-funds the “Rhythms and resonances: coming into contact with otherness” workshop. © Gerd Altmann from Pixabay Interdisciplinary doctoral workshopThe “Rhythms and resonances: coming into contact with otherness”...
